Software available at this site
What this page is
This page provides information about and downloads for methodology for Chromosome Painting. It is not a facility to analyse your genome. Sorry if you were misled by the punchy name!About Chromosome Painting
Painting is an efficient way of identifying important haplotype information from dense genotype data. It describes ancestry in an efficient way suitable for a range of further analyses, including population identification and admixture dating.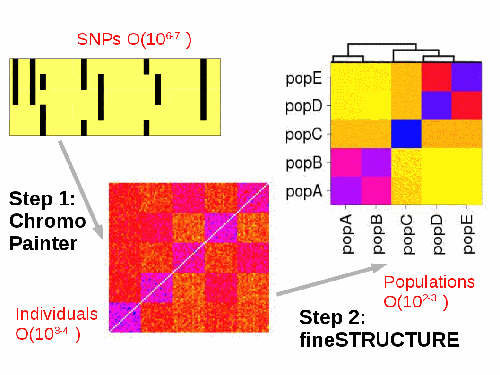
The included ChromoPainter tool finds `haplotypes' in sequence data. Each individual is "painted" as a combination of all other sequences. ChromoPainter can output a range of features, including:
- Sample haplotypes
- Expectations of the number of recombination events at all sites
- A wide range of related features
About fineSTRUCTURE
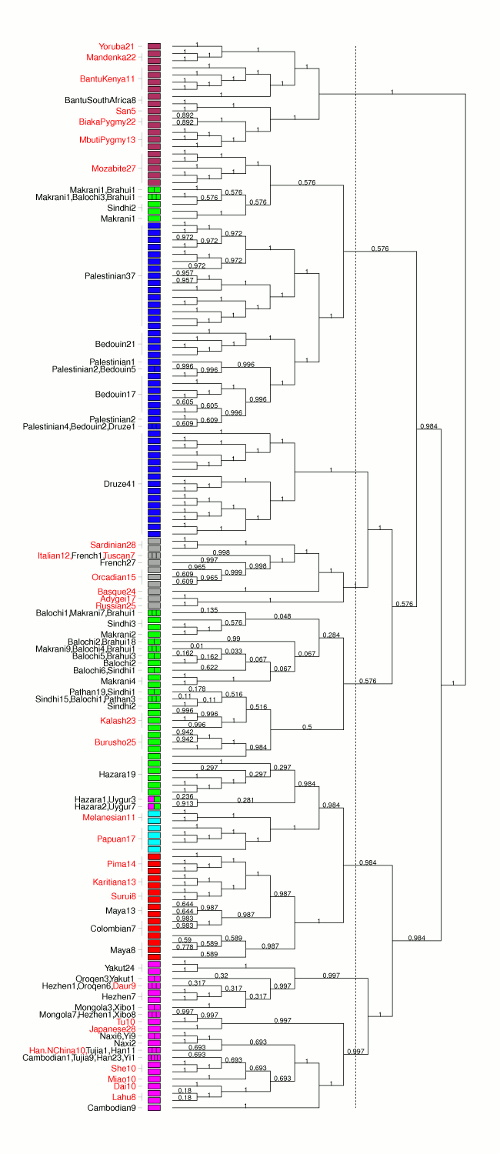
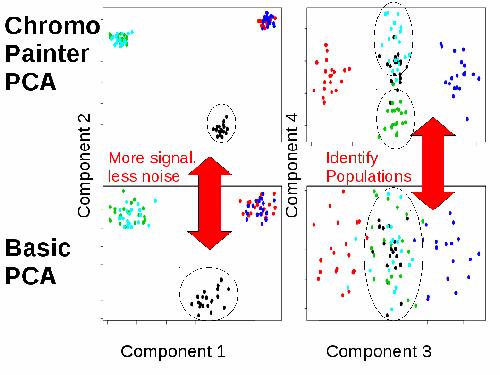
fineSTRUCTURE is a fast and powerful algorithm for identifying population structure using dense sequencing data. By using the output of ChromoPainter as a (nearly) sufficient summary statistic, it is able to perform model-based Bayesian clustering on large datasets, including full resequencing data, and can handle up to 1000s of individuals. Full assignment uncertainty is given.